Rapid Environmental Hygiene Testing Technologies: A Closer Look
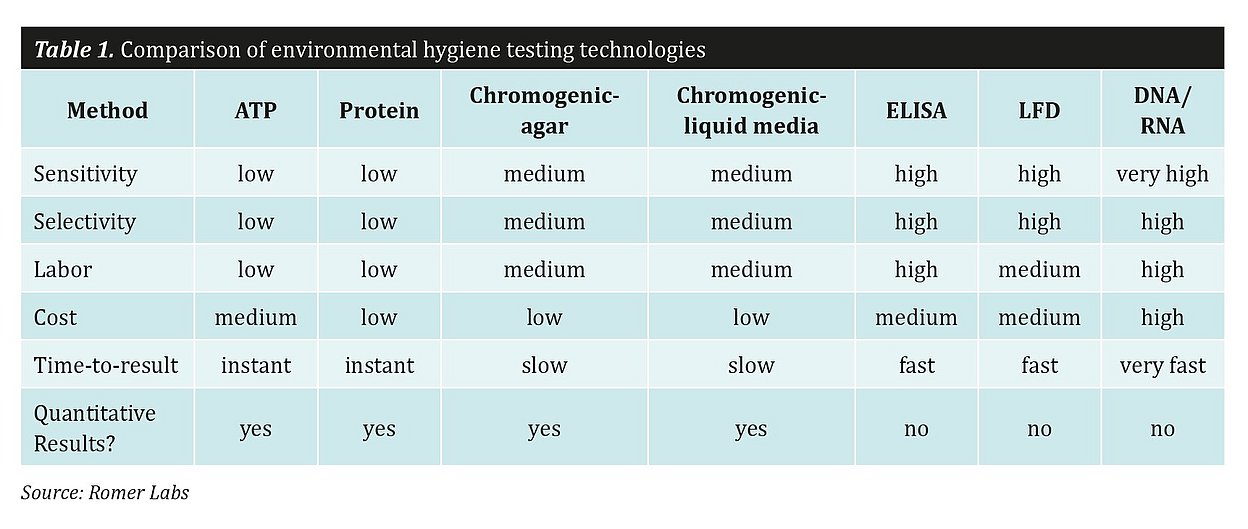
Commonly used environmental testing technologies break down into two general approaches: testing for residues and testing for microorganisms. Here, we investigate the different technologies within each approach, how they work, and their applicability and advantages.
Residue Testing
ATP methods
Adenosine triphosphate (ATP) is a nucleotide used in cells as a coenzyme for delivering energy. It can be thought of as the molecular “unit of currency" for transferring energy within all living cells. Energy is needed for all cellular activities including the synthesis of proteins and membranes, cell movement and cellular division. Energy is transferred when ATP breaks down into adenosine diphosphate and adenosine monophosphate. Hydrolyzing the covalent links of the phosphates liberates energy that is used for reactions.
Commercial ATP test systems harness the luciferin/ luciferase reaction, which is very common in nature, to generate visible light with the energy provided by ATP. The more light is emitted, the more ATP is present, which indicates more food residues or more microorganisms. Yet there is one important caveat: as these systems are widely used for cleaning efficiency validation, disinfectants are also commonly involved in the reaction. These disinfectants can disturb the cell walls of the microorganisms but preserve their ATP, meaning that there may not be a real correlation between living organisms present on the surface and the results of the ATP measurement.
ATP methods harbor a further potential disadvantage: they vary in their applicability depending on the food residue to be detected. For example, ATP testing would not lend itself to testing wheat flour as it is a highly processed matrix that leaves behind little ATP in its residue. Meat product residue, however, contains high levels of ATP.
Total protein detection methods
As another residue testing approach, total protein detection methods test not for microorganisms themselves but rather for amino acids, peptides and proteins. These tests are very fast and deliver results within a minute. The detection is not sensitive enough to detect proteins from single-celled microorganisms. Therefore, a negative result with this type of test does not indicate an absence of the microorganism. Additionally, there is no way to detect specific pathogens using this method. Yet protein detection test kits have their use as they can give indications about cleaning efficiency in a fast and
affordable way.
Microorganism Testing
Chromogenic cultural methods
These are the traditional methods for monitoring the hygiene of the processing environment. Cultural methods are addressed in ISO 18593, “Microbiology of the food chain – Horizontal methods for surface sampling.” Non-selective and selective media can be used to detect specific organisms.
Chromogenic Agar-based methods
Generally, there are two ways to conduct agar-based methods; either directly or indirectly via a dilution step. An advantage of both is that they provide quantitative results.
In direct methods, the agar of plates or dip-slides is pressed directly onto the surface being sampled. Contact plates have the shape of classic Petri dishes and typically have a surface area of 25 cm². Dip-slides are double-sided agar paddles protected by a plastic tube and have an agar surface area of 7-10 cm² on each side of the paddle, which amounts to 14-20 cm². These systems require no additional equipment and the sampling procedure is very fast. However, the sampling area using this method is limited.
Swabs, tissues and sponges are tools for indirect sampling. After they are swabbed over the surface to be tested, they are then diluted in a buffer solution, which is then pipetted into traditional Petri dishes and streaked out. The tested surface area can be much larger and tight spaces and gaps can be tested with an indirect method. However, indirect methods have a downside: there are more handling steps and additional supplies are needed. For pathogen detection, the ISO standard recommends a surface area between 1000 cm² and 3000 cm²; an area this large can only be handled with sponges or swab cloths.
Chromogenic liquid media-based detection methods
This cultural method is only used to detect but not to count specific organisms or organism groups because it does not deliver quantitative results. In liquid media- based methods, swabs are used to take samples and are then placed into tubes filled with selective media. On average, incubation takes at least 48 hours for presumptive negative results. Positive samples are identified by a color change or florescence at a specific wavelength of the enriched sample. These tests are generally easy-to-use and affordable.
The disadvantage of liquid media-based systems is their low degree of sensitivity, a result of highly selective media, which intentionally suppress the growth of other bacteria. There is a certain risk here: bacteria in the processing environment are already stressed and can die if enrichment media are too selective. On the other hand, if the selectivity is too low, it can be difficult to detect specific microorganisms using selective liquid media alone. The interpretation of results can become difficult and subjective, as the color change or florescence is not always strong enough to indicate a definitive result. The main advantage of liquid media-based methods is that they are easy-to-use and affordable.
DNA-based methods
There are three different DNA-based methods: isothermal amplification of DNA, real-time and traditional PCR. Real-time PCR is an established method for the detection of DNA or RNA, while isothermal amplification is a more novel technology. There are clear advantages to using isothermal amplification over real-time PCR: for example, isothermal amplification requires no thermal cycler as the reaction takes place at a constant temperature. The most cost-effective but also most labor-intense DNA-based method is traditional PCR, because it is necessary to perform a hybridization step or to use a rather unspecific dye after amplification to detect amplified DNA or RNA.
In all methods, specific parts of the DNA or RNA are recognized by specific primers and then multiplied by the enzyme polymerase. The sensitivity of these methods is remarkably high, and target regions in the DNA or RNA are well known. The biggest limitation of this system is that it is an enzyme-based system. Enzymes need specified buffer solutions to work properly. Ingredients in disinfectants can influence enzyme activity, which can lead to false negative results. The technology also requires multiple micro-pipetting steps, which can be a serious source of error.
The greatest advantage of DNA-based methods is that non-selective media can be used for the enrichment step thanks to the high sensitivity of the method. Non-selective media are affordable, available from various suppliers and require the lowest biosafety level (level 1). It is also possible to detect low amounts of pathogens on environmental surfaces without enrichment with DNA-based methods, but the sensitivity is low compared to that of enrichment-based methods. Lastly, false positive results from DNA fragments from pathogens that have already been killed could be a major problem.
Immunological methods
Immunoassays are systems that use antibodies to detect the presence of microorganisms in solutions. These antibodies can bind to antigens, such as lipopolysaccharides on the cell surface or flagella of particular microorganisms. After a single or multiple enrichment procedure that includes selective media, pathogens such as Listeria cells are detected with specific antibody- based testing systems.
ELISA (enzyme-linked immunosorbent assay) tests were the first to use immunological technology to detect specific microorganisms. ELISA allows for the quantification of the detected analyte, but the necessity of including an enrichment step precludes any calculation of the initial concentration. Only trained staff can perform the multiple transfer and wash steps that ELISAs require. The development of lateral flow devices, also known as strip tests, solved this problem, rendering the immunological detection procedure much easier and quicker and eliminating the workload of ELISA tests. LFDs with selective antibodies used in combination with high-performing enrichment media enable fast and accurate results without the need for expensive equipment or costs related to training.
Conclusion
Choosing the right hygiene testing technology is not always an easy task. There are several practical considerations that will influence the decision: how many sampling sites must be tested and how important is testing throughput? How can testers optimize time-to-result, given that testing for some pathogenic bacteria is not conducted every day and is more a monitoring task than a quality control procedure? Are quantitative results needed, or is presence/absence testing sufficient? As it is still not possible to get bacterial counts immediately after cleaning, producers may rely on ATP or protein-based testing systems, which can serve as a vague indicator of whether it is safe to begin production, i.e. whether ATP or protein is present.
Also, national regulations can influence this decision. Enrichment-based DNA or immunological methods are superior to other methods in terms of sensitivity and selectivity and should be the systems of choice for pathogen detection. For detecting and counting indicator organisms or monitoring general hygiene, more affordable systems such as dip-slides or ATP systems may be used.
Published on:
Microbiology